Our Research
Research in the Hiller lab revolves around a central question in genomics and evolutionary biology:
What is the genomic basis of phenotypic differences between species?
Our focus is explicitly on differences between species as opposed to differences within a species. We aim at discovering the genomic determinants of macroevolutionary trait differences, which is important to understand how nature’s spectacular phenotypic diversity has evolved. Our current focus is on studying the genomic basis of vertebrate traits that have potential applications in biomedicine or other fields.
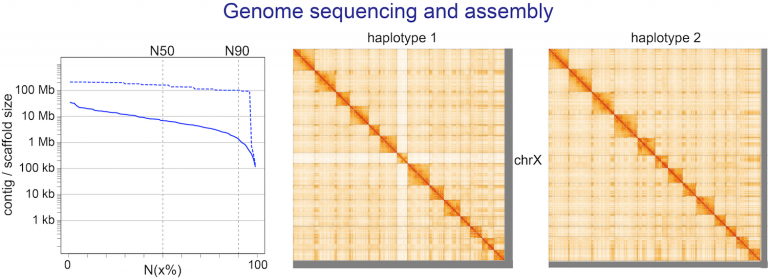
- sequencing reference-quality genomes and transcriptomes of selected species with interesting traits,
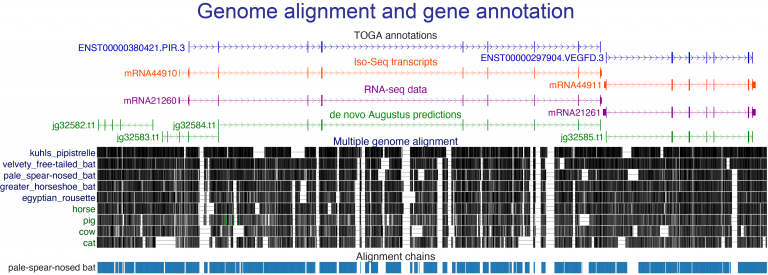
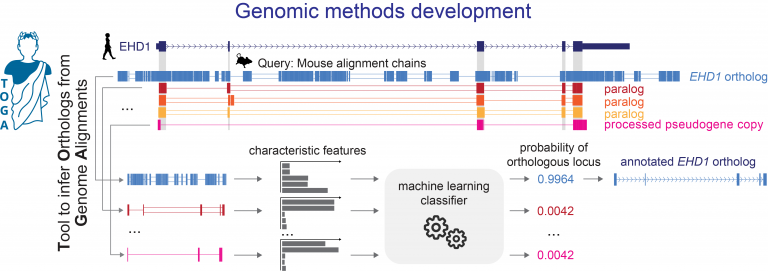
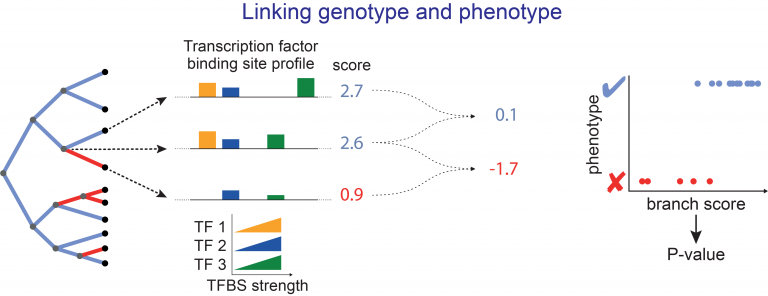
- developing new computational approaches to align genes and genomes, to annotate genes and infer orthologs, to accurately detect relevant evolutionary changes in functional genomic regions, and to discover associations between genotype and phenotype,
- running large-scale comparative genomic screens using existing and newly-sequenced genomes and our powerful methods toolbox to reveal the genomic basis of phenotypic adaptations and link phenotypic to genomic differences.
More information about our current projects:
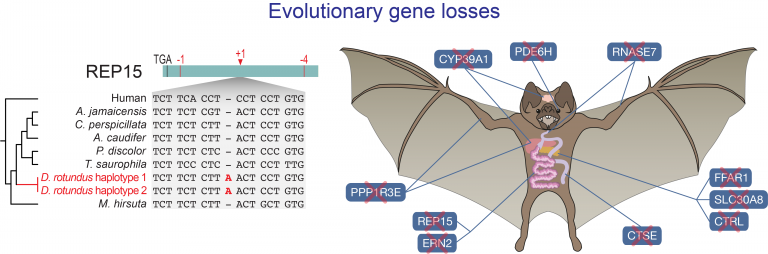
- Genomic underpinnings of exceptional traits in bats
- Genomics of dietary adaptations
- Adaptive gene losses
- Forward Genomics: Linking phenotypic to genomic changes across species
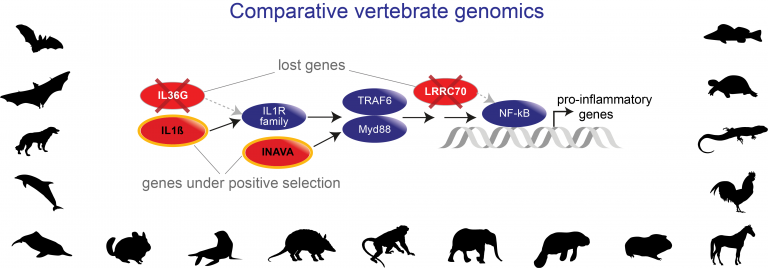
- Development of comparative genomics methods
Videos
- Forward Genomics and linking phenotype to genotype (ESEB 2019; 10 min)
- Genomic determinants of adaptations in bats (PAG 2020; 26 min)
- Hat Evolution wirklich stattgefunden? (YouTube in German, 2021)
- Fledermäuse: Flug, Viren und ihr Genom erklärt (YouTube in German, 2026)
Selected publications
- Osipova E, Ko MC, Petricek KM, Sin SYW, Brown T, Winkler S, Pippel M, Jarrells J, Weiche S, Mosbech MB, Taborsak-Lines F, Wang C, Contreras-Lopez O, Olsen RA, Ewels P, Mendez-Aranda D, Gaede A, Sadanandan K, Low GW, Monte A, Ballerstaedt N, Adreani NM, Mentesana L, von Bayern A, Rico-Guevara A, Edwards SV, Frankl-Vilches C, Kuhl H, Bakker A, Gahr M, Altshuler DL, Buttemer WA, Schupp M, Baldwin MW #, Hiller M #, Sackton TB #. Convergent and lineage-specific genomic changes shape adaptations in sugar-consuming birds. Science, 391(6788), 2026
- Morales AE, Dong Y, Brown T, Baid K, Kontopoulos DK, Gonzalez V, Huang Z, Ahmed AW, Bhuinya A, Hilgers L, Winkler S, Hughes G, Li X, Lu P, Yang Y, Kirilenko BM, Devanna P, Lama TM, Nissan Y, Pippel M, Dávalos LM, Vernes SC, Puechmaille SJ, Rossiter SJ, Yossi Y, Prescott JB, Kurth A, Ray DA, Lim BK, Myers E, Teeling EC, Banerjee A, Irving AT #, Hiller M #. Bat genomes illuminate adaptations to viral tolerance and disease resistance. Nature, 637, 449–458, 2025
- Kirilenko BM, Munegowda C, Osipova E, Jebb D, Sharma V, Blumer M, Morales AE, Ahmed AW, Kontopoulos DG, Hilgers L, Lindblad-Toh K, Karlsson EK, Zoonomia Consortium, Hiller M #. Integrating gene annotation with orthology inference at scale. Science, 380 (6643), eabn3107, 2023
- Osipova E, Barsacchi R, Brown T, Sadanandan K, Gaede AH, Monte A, Jarrells J, Moebius C, Pippel M, Altshuler DL, Winkler S, Bickle M, Baldwin MW, Hiller M #. Loss of a gluconeogenic muscle enzyme contributed to adaptive metabolic traits in hummingbirds. Science, 379(6628), 185-190, 2023 pdf of article
- Blumer M, Brown T, Freitas MB, Destro AL, Oliveira JA, Morales A, Schell T, Greve C, Pippel M, Jebb D, Hecker N, Ahmed AW, Kirilenko BM, Foote M, Janke A, Lim BK, Hiller M #. Gene losses in the common vampire bat illuminate molecular adaptations to blood feeding. Science Advances, 8 (12), eabm6494, 2022
press and blogs are listed at altmetric - Jebb D, Huang Z, Pippel M, Hughes GM, Lavrichenko K, Devanna P, Winkler S, Jermiin LS, Skirmuntt EC, Katzourakis A, Burkitt-Gray L, Ray DA, Sullivan KAM, Roscito JG, Kirilenko BM, Dávalos LM, Corthals AP, Power ML, Jones G, Ransome RD, Dechmann D, Locatelli AG, Puechmaille SJ, Fedrigo O, Jarvis ED, Hiller M #, Vernes SC #, Myers EW #, Teeling EC #. Six reference-quality genomes reveal evolution of bat adaptations. Nature, 583, 578–584, 2020
See also Nature Reviews Genetics highlight and Science feature. More press. - Huelsmann M, Hecker N, Springer MS, Gatesy J, Sharma V, Hiller M #. Genes lost during the transition from land to water in cetaceans highlight genomic changes associated with aquatic adaptations. Science Advances, 5(9), eaaw6671, 2019
some press: New York Times, Science News, Smithsonian, F1000 recommended - Roscito JG, Sameith K, Parra G, Langer BE, Petzold A, Moebius C, Bickle M, Rodrigues MT, Hiller M #. Phenotype loss is associated with widespread divergence of the gene regulatory landscape in evolution. Nature Communications, 9:4737, 2018
- Sharma V, Hecker N, Roscito JG, Foerster L, Langer BE, and Hiller M #. A genomics approach reveals insights into the importance of gene losses for mammalian adaptations. Nature Communications, 9(1), 1215, 2018
- Prudent X, Parra G, Schwede P, Roscito JG, Hiller M #. Controlling for phylogenetic relatedness and evolutionary rates improves the discovery of associations between species’ phenotypic and genomic differences. Mol Biol Evol, 33(8), 2135-5, 2016.
See also MBE News. - Hiller M #, Schaar BT, Indjeian VB, Kingsley DM, Hagey LR, and Bejerano G #. A “forward genomics” approach links genotype to phenotype using independent phenotypic losses among related species. Cell Reports, 2(4), 817-823, 2012. Read more on F1000
News
03/2026 Michael wrote a Nature Methods News & Views on deep learning based gene finders such as ANNEVO.
02/2026 Happy to share that Katya’s work on convergent and lineage-specific genomic adaptations in sugar-feeding birds is published in Science.
07/2025 Leon’s review on linking phenotype to genotype using comprehensive
genomic comparisons is out.
06/2025 The Cetacean genome infrastructure paper that we contributed to is out in Frontiers Marine Science.
06/2025 We welcome our new PhD student Ariel Eraso.
04/2025 Mridula Nandakumar, new postdoc, and Lucas Canesin, new Bioinformatician, started in the lab. Welcome.
02/2025 Bernhard’s work on long read sequencing of ethanol and difficult-to-sequence samples is out in Genome Biology.
02/2025 Leon’s work how to avoid a common PSMC artifact is out in Current Biology. Congrats !!
01/2025 Happy to see work from Ariadna, Dimitris and other lab members on immune system adaptations in bats out in Nature.
01/2025 Dimitris’ publication on the evolution of daily torpor and hibernation in mammals and birds is out in Functional Ecology.
01/2025 We welcome our new PhD student Michele Albertini.
10/2024 Happy to see our Garden Dormouse genome study with implications for its conservation out in Genome Research.
10/2024
We are have a Postdoc and Bioinformatician position to join our BatProtect project. 6 year funding available.
09/2024
Welcome to our new PhD student Alejandro Gonzales, who will work on gene annotation methods.
06/2024
Welcome to our new postdoc Xueling Yi, who will work on dietary adaptations in bats,
10/2023
Michael has been awarded an ERC Synergy grant ‘BatProtect‘ to uncover the molecular mechanisms behind longevity and disease resistance in bats with three other PIs.
10/2023
Leon was awarded a well-funded DFG grant to investigate genome evolution and population collapse in eels. Congrats !!
04/2023
Huge congrats to Bogdan and the lab for the TOGA publication, now out in Science
02/2023
Superathlete vs. couch potato: Scientific American featured our hummingbird story together with Nick Rohner’s cavefish work.
01/2023
Katya’s work on FBP2 loss in ancestral hummingbirds and metabolic muscle adaptations for hovering flight is now published in Science.
10/2022
Great to see 2 graduated PhD students moving on. Ekaterina does a postdoc at Harvard. Bogdan is a scientist at Quantgene (genomic medicine). All the best!!
08/2022
Great to see Henrike’s paper on predicting novel eye genes based on evolutionary gene loss signatures out in Elife.
07/2022
Welcome to our new PhD student Yury Malovichko.
06/2022
Attending EuroEvoDevo in Naples with 3 talks by lab members.
05/2022
Huge congrats to Ekaterina Osipova, who did a great job defending her PhD.
04/2022
We welcome Bernhard Bein as a new PhD student in the lab.
04/2022
Welcome to our new Postdoc Shenglin Liu !!
03/2022
We contributed a high-quality gene annotation (BUSCO 99.4%) for the helmeted honeyeater.
03/2022
Our vampire bat gene loss paper is out in Science Advances. Some press
02/2022
Big congrats to Dimitris Kontopoulos for winning an EMBO Postdoc Fellowship.
01/2022
Juliana’s paper on convergent limb losses in reptiles is published in Cell Reports.
01/2022
Our assembly gap closing method DENTIST is published at GigaScience.
12/2021
LOEWE-TBG successfully got funding for the next 3 years !!
11/2021
Huge congrats to Bogdan Kirilenko, our TOGA master, for defending his PhD.
10/2021
Excellent blog on our preprint on gene losses
in vampire bats.
09/2021
Ekaterina Osipova gave a great talk @EMBOEvolAnimalGenomes on gene losses in hummingbirds and metabolic adaptation
08/2021
Welcome to our new Postdoc Leon Hilgers !!
05/2021
Lab move to TBG and Senckenberg in Frankfurt